Abstract: A collaborative team including Kanazawa University used force spectroscopy to determine the transmembrane structure of the protein prestin, which is an important motor protein of the inner ear. They functionalized the protein with biotin and were then able to achieve single protein interaction with an atomic force microscopy cantilever modified with streptavidin. The strength of the bond was sufficient to allow stretching of the protein, which suggested a structure with 12 transmembrane domains.
Kanazawa, Japan – As mammals we have very sensitive hearing thanks to a process known as cochlear amplification. This amplification is down to sensory cells in the inner ear called outer hair cells (OHCs), and it is thought to depend on the way a protein called prestin—found in the OHC membrane—responds to changes in electric potential. However, the membrane structure of prestin is unknown, so a team including a researcher from Kanazawa University has used force spectroscopy to determine it. Their findings are published in Journal of Biomechanical Science and Engineering.
Proteins are long chain molecules made of smaller units called amino acids. These long chains fold up on themselves to give 3D structures, bringing parts of the protein closer together leading to functional properties. Prestin is known to have 744 amino acids, but its 3D structure in the OHC membrane is unknown.
Determining membrane structures requires high resolution techniques, which often require a lot of protein. An alternative approach is to analyze what happens when a protein molecule is stretched. Single molecule force spectroscopy (SMFS) uses a tiny arm—less than 0.05 mm across—to pull a protein from the membrane.
However, biological systems contain numerous proteins, so identifying and anchoring the protein of interest is crucial. Previous attempts to isolate prestin for SMFS have had limitations. The researchers therefore devised a technique that uses a strong and highly specific interaction—widely used in biochemistry—between the molecule biotin and the protein streptavidin.
“The beauty of adding biotin to prestin is that it is small, so it doesn’t have a significant effect on the way prestin would otherwise behave,” explains Associate Professor Michio Murakoshi, study first author. “The biotin interacts with streptavidin that we apply to the arm, and the interaction is strong enough to hold while the protein is pulled from the membrane.”
The researchers acquired force-extension curves, showing how much force was needed to extract the protein from the plasma membrane. The curves had a saw-toothed pattern—because the force needed increased sharply at certain points in the extension when the protein was unfolding. A bit like a knotted string snagging at sticking points when trying to pull it through a tangle.
“Our results suggest that prestin has 12 transmembrane domains, which supports the predictions of a previous model,” says Murakoshi. “Our findings demonstrate the potential of our biotin-streptavidin method for investigating proteins using SMFS, and provide important information on the response of OHC membrane proteins, contributing to a better understanding of cochlear amplification.”
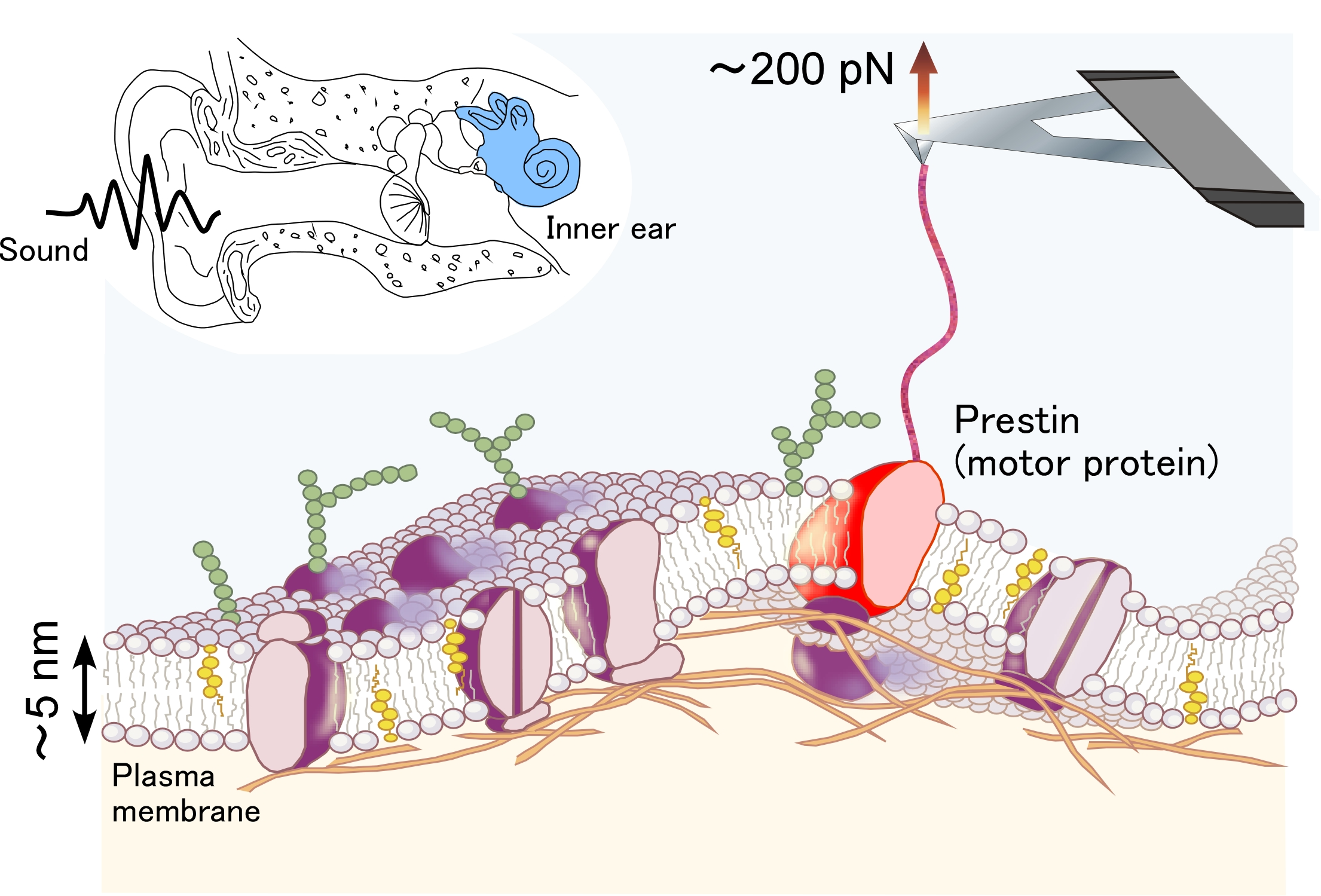
Figure 1.
Force spectroscopy of prestin, the motor protein in the inner ear.
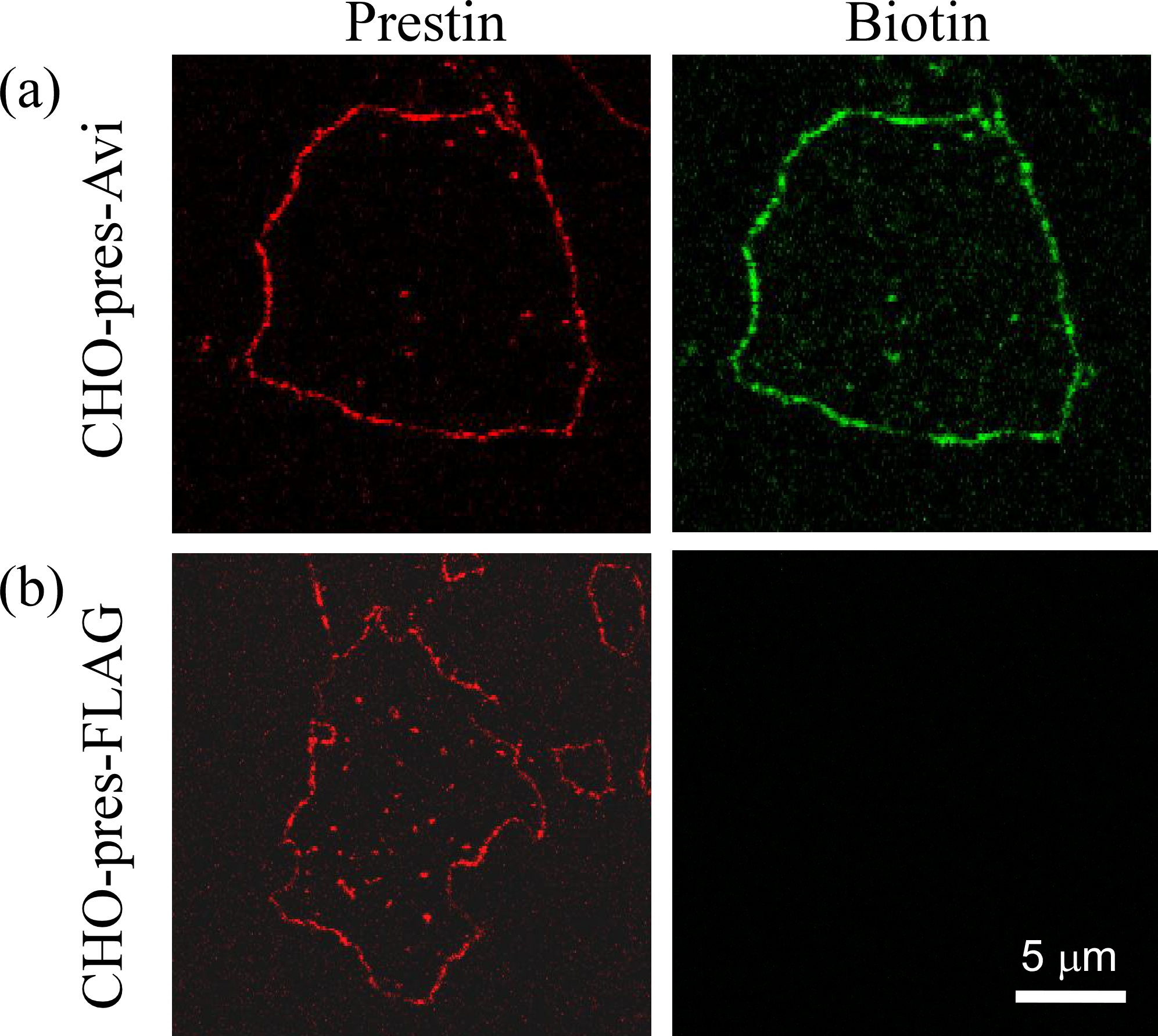
Figure 2
Fluorescence images of the isolated plasma membrane of CHO cells. (a) Avi-tagged prestin (CHO-pres-Avi). (b) FLAG-tagged prestin (CHO-pres-FLAG). Prestin labeling (red) was observed in both cell lines (left panels). Biotin labeling (green) was only confirmed in CHO-pres-Avi.
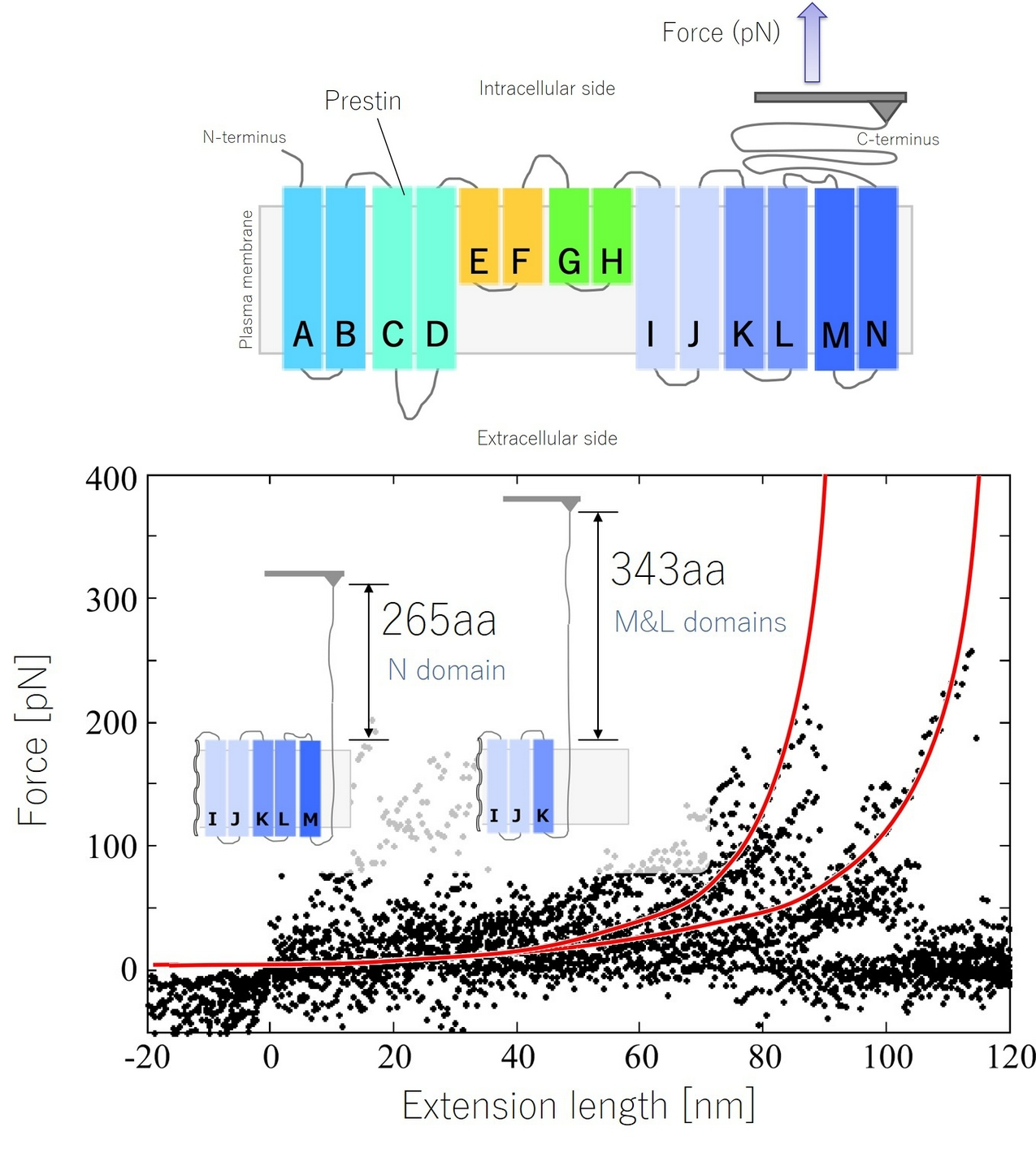
Figure 3
Structural analysis of prestin by force spectroscopy using an AFM. Force-extension (FE) curve obtained from the N-M domain.
[Article]
Title: Analysis of membrane structure of the inner ear motor protein prestin by force spectroscopy
Journal: Journal of Biomechanical Science and Engineering
Authors: Michio Murakoshi and Hiroshi Wada



 PAGE TOP
PAGE TOP